-Search query
-Search result
Showing 1 - 50 of 142 items for (author: miyazaki & n)

EMDB-16440: 
C1 reconstruction of Tanay virus particle
Method: single particle / : Okamoto K, Song C, Miyazaki N, Murata K
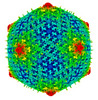

EMDB-32974: 
Capsid structure of Staphylococcus jumbo bacteriophage S6
Method: single particle / : Koibuchi W, Uchiyama J, Matsuzaki S, Murata K, Iwasaki K, Miyazaki N

PDB-7x30: 
Capsid structure of Staphylococcus jumbo bacteriophage S6
Method: single particle / : Koibuchi W, Uchiyama J, Matsuzaki S, Murata K, Iwasaki K, Miyazaki N

EMDB-35535: 
Human ornithine transcarbamylase expressed using mRNA lipid nanoparticles in human cells and PEGylated
Method: single particle / : Kondo K, Yamazaki K, Kubara K, Ishii S, Suzuki Y, Miyazaki T, Mitsuhashi K, Ito M, Tsukahara K
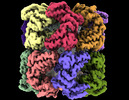
EMDB-33349: 
Cryo-EM structure of GroEL bound to unfolded substrate (UGT1A) at 2.8 Ang. resolution (Consensus Refinement)
Method: single particle / : Stapleton K, Takagi J, Mizohata E
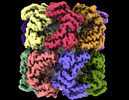
EMDB-33350: 
Cryo-EM structure of double occupied ring (DOR) of GroEL-UGT1A complex at 2.7 Ang. resolution
Method: single particle / : Stapleton K, Takagi J, Mizohata E
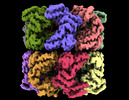
EMDB-33351: 
Cryo-EM structure of single empty ring 2 (SER2) of GroEL-UGT1A complex at 3.2 Ang. resolution
Method: single particle / : Stapleton K, Takagi J, Mizohata E
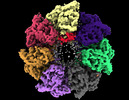
EMDB-33352: 
Cryo-EM structure of occupied ring subunit 4 (OR4) of GroEL complexed with polyalanine model of UGT1A from GroEL-UGT1A double occupied ring complex
Method: single particle / : Stapleton K, Takagi J, Mizohata E

EMDB-33353: 
Cryo-EM structure of empty ring subunit 1 (ER1) from single empty ring of GroEL-UGT1A complex
Method: single particle / : Stapleton K, Takagi J, Mizohata E

EMDB-33354: 
Cryo-EM structure of empty ring subunit 2 (ER2) from GroEL-UGT1A single empty ring complex
Method: single particle / : Stapleton K, Takagi J, Mizohata E
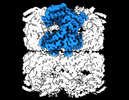
EMDB-33355: 
Cryo-EM structure of occupied ring subunit 1 (OR1) of GroEL from GroEL-UGT1A double occupied ring complex
Method: single particle / : Stapleton K, Takagi J, Mizohata E
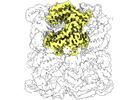
EMDB-33356: 
Cryo-EM structure of occupied ring subunit 2 (OR2) of GroEL from GroEL-UGT1A double occupied ring complex
Method: single particle / : Stapleton K, Takagi J, Mizohata E
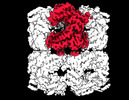
EMDB-33357: 
Cryo-EM structure of occupied ring subunit 3 (OR3) of GroEL from GroEL-UGT1A double occupied ring complex
Method: single particle / : Stapleton K, Takagi J, Mizohata E
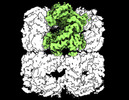
EMDB-33358: 
Cryo-EM structure of occupied ring subunit 4 (OR4) of GroEL from GroEL-UGT1A double occupied ring complex
Method: single particle / : Stapleton K, Takagi J, Mizohata E
EMDB-33007: 
A CBg-ParM filament with ADP
Method: helical / : Koh A, Ali S, Robinson R, Narita A
EMDB-33008: 
A CBg-ParM filament with GTP and a short incubation time
Method: helical / : Koh A, Ali S, Robinson R, Narita A
EMDB-33009: 
A CBg-ParM filament with ADP
Method: helical / : Koh A, Ali S, Robinson R, Narita A
EMDB-33012: 
A CBg-ParM filament with GTP or GDPPi
Method: helical / : Koh A, Ali S, Robinson R, Narita A

EMDB-32614: 
Overall structure of Helicobacter pylori bacteriophage KHP30
Method: single particle / : Kamiya R, Uchiyama J, Matsuzaki S, Murata K, Iwasaki K, Miyazaki N

EMDB-32616: 
Tail structure of Helicobacter pylori bacteriophage KHP30
Method: single particle / : Kamiya R, Uchiyama J, Matsuzaki S, Murata K, Iwasaki K, Miyazaki N

PDB-7wmp: 
Tail structure of Helicobacter pylori bacteriophage KHP30
Method: single particle / : Kamiya R, Uchiyama J, Matsuzaki S, Murata K, Iwasaki K, Miyazaki N

EMDB-15855: 
Rosellinia necatrix megabirnavirus 1-W779 full capsid
Method: single particle / : Wang H, Okamoto K, Miyazaki N, Suzuki N

EMDB-15857: 
Rosellinia necatrix megabirnavirus 1-W779 empty capsid
Method: single particle / : Wang H, Okamoto K, Miyazaki N, Suzuki N

EMDB-15859: 
Rosellinia necatrix megabirnavirus 1-W779 full capsid with Crown protein
Method: single particle / : Wang H, Okamoto K, Miyazaki N, Suzuki N

PDB-8b4z: 
Rosellinia necatrix megabirnavirus 1-W779 full capsid
Method: single particle / : Wang H, Okamoto K, Miyazaki N, Suzuki N

PDB-8b59: 
Rosellinia necatrix megabirnavirus 1-W779 Crown protein
Method: single particle / : Wang H, Okamoto K, Miyazaki N, Suzuki N
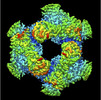
EMDB-32571: 
Cryo-EM structure of GH31 alpha-1,3-glucosidase from Lactococcus lactis subsp. cremoris
Method: single particle / : Ikegaya M, Moriya T, Adachi N, Kawasaki M, Park EY, Miyazaki T

PDB-7wlg: 
Cryo-EM structure of GH31 alpha-1,3-glucosidase from Lactococcus lactis subsp. cremoris
Method: single particle / : Ikegaya M, Moriya T, Adachi N, Kawasaki M, Park EY, Miyazaki T

EMDB-14174: 
TMEM106B filaments with Fold I from Alzheimer's disease (case 1)
Method: helical / : Lovestam S, Schweighauser M, Scheres SHW

EMDB-14176: 
TMEM106B filaments with Fold I-d from Multiple system atrophy (case 18)
Method: helical / : Lovestam S, Schweighauser M, Scheres SHW

EMDB-14187: 
TMEM106B filaments with Fold IIa from Multiple system atrophy (case 19)
Method: helical / : Lovestam S, Schweighauser M, Scheres SHW

EMDB-14188: 
TMEM106B filaments with Fold IIb from Multiple system atrophy (case 19)
Method: helical / : Lovestam S, Schweighauser M, Scheres SHW

EMDB-14189: 
TMEM106B filaments with Fold III from Multiple system atrophy (case 17)
Method: helical / : Lovestam S, Schweighauser M, Scheres SHW
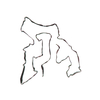
PDB-7qvc: 
TMEM106B filaments with Fold I from Alzheimer's disease (case 1)
Method: helical / : Lovestam S, Schweighauser M, Scheres SHW
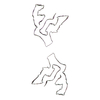
PDB-7qvf: 
TMEM106B filaments with Fold I-d from Multiple system atrophy (case 18)
Method: helical / : Lovestam S, Schweighauser M, Scheres SHW
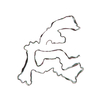
PDB-7qwg: 
TMEM106B filaments with Fold IIa from Multiple system atrophy (case 19)
Method: helical / : Lovestam S, Schweighauser M, Scheres SHW
PDB-7qwl: 
TMEM106B filaments with Fold IIb from Multiple system atrophy (case 19)
Method: helical / : Lovestam S, Schweighauser M, Scheres SHW

PDB-7qwm: 
TMEM106B filaments with Fold III from Multiple system atrophy (case 17)
Method: helical / : Lovestam S, Schweighauser M, Scheres SHW

EMDB-31905: 
Structure of C2S2M2-type PSII-FCPII supercomplex from diatom
Method: single particle / : Nagao R, Kato K, Akita F, Miyazaki N, Shen JR

EMDB-31906: 
Structure of S1M1-type FCPII complex from diatom
Method: single particle / : Nagao R, Kato K, Akita F, Miyazaki N, Shen JR

PDB-7vd5: 
Structure of C2S2M2-type PSII-FCPII supercomplex from diatom
Method: single particle / : Nagao R, Kato K, Akita F, Miyazaki N, Shen JR

PDB-7vd6: 
Structure of S1M1-type FCPII complex from diatom
Method: single particle / : Nagao R, Kato K, Akita F, Miyazaki N, Shen JR

EMDB-30820: 
Structure of Wild-type PSI monomer1 from Cyanophora paradoxa
Method: single particle / : Kato K, Nagao R, Akita F, Miyazaki N, Shen JR

EMDB-30821: 
Structure of Wild-type PSI monomer2 from Cyanophora paradoxa
Method: single particle / : Kato K, Nagao R, Akita F, Miyazaki N, Shen JR

EMDB-30822: 
Structure of Wild-type PSI tetramer from Cyanophora paradoxa
Method: single particle / : Kato K, Nagao R, Akita F, Miyazaki N, Shen JR

EMDB-30823: 
Structure of GraFix PSI tetramer from Cyanophora paradoxa
Method: single particle / : Kato K, Nagao R, Akita F, Miyazaki N, Shen JR

PDB-7dr0: 
Structure of Wild-type PSI monomer1 from Cyanophora paradoxa
Method: single particle / : Kato K, Nagao R, Akita F, Miyazaki N, Shen JR

PDB-7dr1: 
Structure of Wild-type PSI monomer2 from Cyanophora paradoxa
Method: single particle / : Kato K, Nagao R, Akita F, Miyazaki N, Shen JR

PDB-7dr2: 
Structure of GraFix PSI tetramer from Cyanophora paradoxa
Method: single particle / : Kato K, Nagao R, Akita F, Miyazaki N, Shen JR

EMDB-30793: 
Capsid structure of human sapovirus
Method: single particle / : Miyazaki N, Murakami K, Oka T, Iwasaki K, Katayama K, Murata K
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
